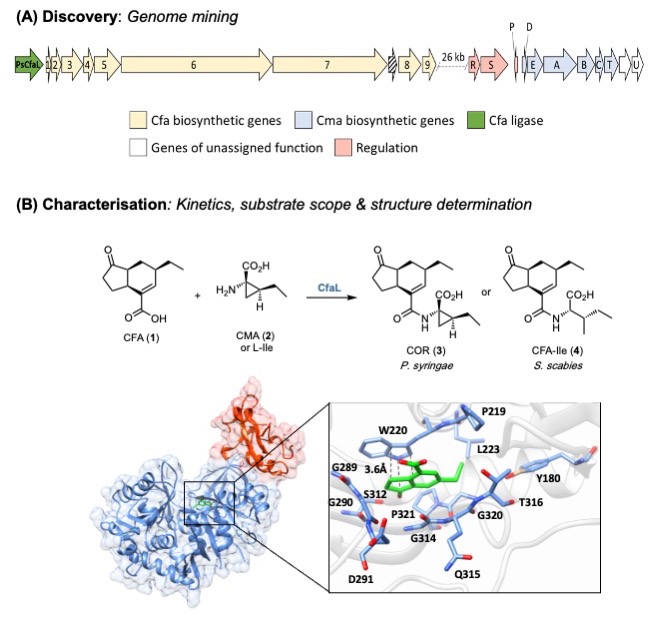
We are interested in the discovery, characterisation and engineering of biosynthetic pathways to nonribosomal peptides, polyketides and other bioactive natural products. More sustainable routes to the next generation of antibiotics to combat antimicrobial resistance (AMR) and treated neglected disease is the main focus. In addition, we have been working on pathways that produce natural products for agrochemical applications, which can help boost crop yields to feed the growing population. For example, we have characterised key ligase enzymes (CfaL) in the biosynthesis of coronatine and related bacterial phytotoxins that mimic the hormone jasmonyl-l-isoleucine ( JA-Ile), which mediates physiologically important plants [1]. Coronatine-like phytotoxins disrupt these essential pathways have potential in the development of safer, more selective herbicides.
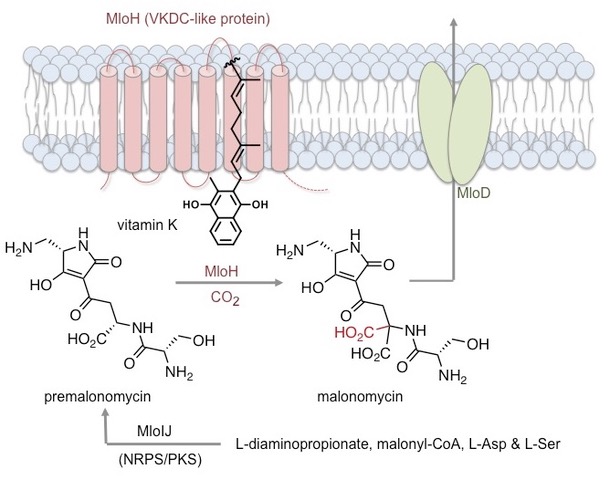
Most recently we discovered a new biosynthetic pathway, including a hybrid nonribosomal peptide synthetase (NRPS)-polyketide synthase (PKS) assembly line and a novel carboxylase enzyme (MloH), that produces a structurally unique antibiotic malonomycin [2]. This study, provides the first example of a vitamin-K dependent carboxylase (VKDC) enzyme in secondary metabolism and the first evidence for the function of VKDC-like proteins in prokaryotes. An interdisciplinary approach combining genome sequencing, bioinformatics, CRISPR-Cas9 gene editing, in vitro enzyme assays, 18O labeling, synthetic and analytical chemistry was used to elucidate the biosynthesis of malonomycin. The study also showed that many VKDC-like enzymes are present in bacteria, opening the way for the discovery of new pathways and novel antibiotics that are urgently required to combat emerging drug-resistant pathogens [2]. We also helped characterise the NRPS-PKS assembly line that delivers the epoxyketone proteasome inhibitor TMC-86A, which could be used to generate new antitumor agents [3].
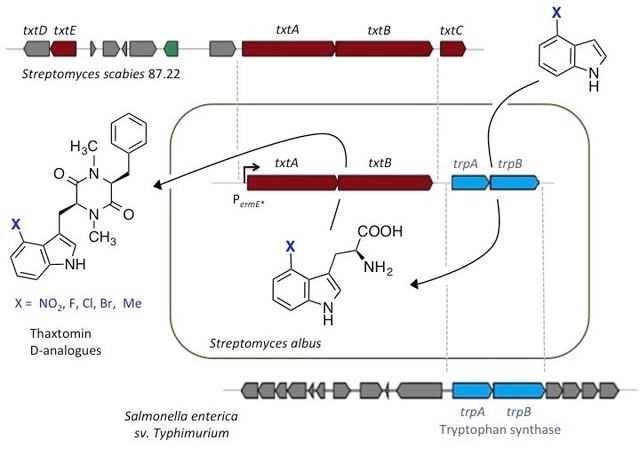
Our lab has also recently developed new synthetic biology approaches to deliver improved natural product variants. For example, genes encoding two NRPS and a promiscuous tryptophan synthase from different species, were assembled in a heterologous host to create a de novo pathway to “non‐natural” thaxtomin phytotoxins, with improved stability, that are also of industrial interest as herbicides for crop protection [4].
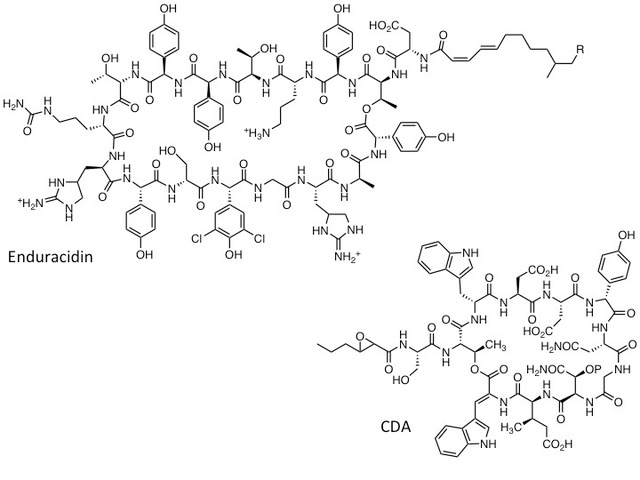
Earlier work in our lab focused on elucidating the biosynthetic origins of the calcium dependant antibiotics (CDA) from Streptomyces coelicolor, which are lipopeptides similar to daptomycin [5,6,7]. We went on to develop a wide range of biosynthetic engineering approaches (combinatorial biosynthesis) which enabled us to generate many “non-natural” lipopeptide variants by altering the specificity of the biosynthetic enzymes [7,8,9]. We have also developed methods for engineering the biosynthesis of the lipopeptide antibiotic enduracidin, using a membrane associated polyprenyl phosphomannose-dependent glycosyltransferases from ramoplanin biosynthesis to deliver novel lipoglycopeptide antibiotics [10].
[1] M. Winn, F. Wang, M. Rowlinson, L. Bering, D. Francis, C. Levy & J Micklefield Nature 2021, 593, 391–398; [2] B. J. C. Law, Y. Zhuo, D. Francis, M. Winn, Y. Zhang, M. Samborskyy, A. Murphy, L. Ren, P. F. Leadlay & J. Micklefield. Nature Catalysis 2018, 1, 977–984; [3] D. Zabala; J. W. Cartwright; D. M. Roberts; B. J. C. Law; L. Song; M. Samborskyy; P. F. Leadlay; J. Micklefield; G. L. Challis. J. Am. Chem. Soc. 2016, 138, 4342–4345; [4] M. Winn, D. Francis & J. Micklefield. Angew. Chem. Int. Ed. 2018, 57, 6830–6833; [5] C. Milne, A. Powell, J. Jim, M. Al Nakeeb C. P. Smith & J. Micklefield. J. Am. Chem. Soc. 2006, 128, 11250-11259; [6] C. Mahlert, F. Kopp, J. Thirlway, J. Micklefield & M. A. Marahiel. J. Am. Chem. Soc. 2007, 129, 12011-12018; [7] A. Powell, M. Borg B. Amir-Heidari, J. M. Neary, J. Thirlway, B. Wilkinson, C.P. Smith and J. Micklefield. J. Am. Chem. Soc. 2007, 129, 15182-15191; [8] J. Thirlway, R. Lewis, L. Nunns, M. Al Nakeeb, M. Styles, A.-W. Struck, C. P. Smith & J. Micklefield. Angew. Chem. Int. Ed. 2012, 51, 7181–7184; [9] R. Lewis, L. Nunns, J. Thirlway, K. Carroll, C. P. Smith, J. Micklefield. Chem. Commun. 2011, 47, 1860-1862; [10] M.-C. Wu, M. Q. Styles, B. J. C. Law, A. W. Struck, L. Nunns and J. Micklefield. Microbiology 2015, 161, 1338-1347.